
Chinese Journal OF Rice Science ›› 2015, Vol. 29 ›› Issue (3): 241-249.DOI: 10.3969/j.issn.1001G7216.2015.03.003
• ResearchPaper • Previous Articles Next Articles
Qing DONG1,2, Ying-xin ZHANG1,2, Zhen-hua ZHANG1,2, Quan ZHOU1,2, Ya-zhi QIN1,2,3, Hong WANG1,2, Shi-hua CHENG1,2, Li-yong CAO1,2,*( ), Xi-hong SHEN1,2,*(
), Xi-hong SHEN1,2,*( )
)
Received:2014-11-27
Revised:2014-12-24
Online:2015-05-10
Published:2015-05-10
Contact:
Li-yong CAO, Xi-hong SHEN
董青1,2, 张迎信1,2, 张振华1,2, 周全1,2, 秦亚芝1,2,3, 王宏1,2, 程式华1,2, 曹立勇1,2,*( ), 沈希宏1,2,*(
), 沈希宏1,2,*( )
)
通讯作者:
曹立勇,沈希宏
基金资助:CLC Number:
Qing DONG, Ying-xin ZHANG, Zhen-hua ZHANG, Quan ZHOU, Ya-zhi QIN, Hong WANG, Shi-hua CHENG, Li-yong CAO, Xi-hong SHEN. Genetic Analysis and Gene Mapping of a Yellow-green Leaf Mutant w390 in Rice (Oryza sativa L.)[J]. Chinese Journal OF Rice Science, 2015, 29(3): 241-249.
董青, 张迎信, 张振华, 周全, 秦亚芝, 王宏, 程式华, 曹立勇, 沈希宏. 水稻黄绿叶突变体w390的遗传分析和基因定位[J]. 中国水稻科学, 2015, 29(3): 241-249.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.3969/j.issn.1001G7216.2015.03.003
| 光合色素含量 Photosynthetic pigment content | 中恢8015 R8015 | w390 | P |
|---|---|---|---|
| 叶绿素a Chlorophyll a | 2.65±0.03 | 1.34±0.05 | <0.0001 |
| 叶绿素b Chlorophyll b | 0.74±0.05 | 0.00 | <0.0001 |
| 类胡萝卜素 Carotenoids | 0.66±0.04 | 0.36±0.01 | <0.0003 |
| 总含量 Total content | 4.05±0.07 | 1.70±0.06 | <0.0001 |
Table 1 Comparison of photosynthetic pigment contents in leaves between the mutant and its wild-type parent.mg/g
| 光合色素含量 Photosynthetic pigment content | 中恢8015 R8015 | w390 | P |
|---|---|---|---|
| 叶绿素a Chlorophyll a | 2.65±0.03 | 1.34±0.05 | <0.0001 |
| 叶绿素b Chlorophyll b | 0.74±0.05 | 0.00 | <0.0001 |
| 类胡萝卜素 Carotenoids | 0.66±0.04 | 0.36±0.01 | <0.0003 |
| 总含量 Total content | 4.05±0.07 | 1.70±0.06 | <0.0001 |
| 性状 Trait | 中恢8015 R8015 | w390 | P |
|---|---|---|---|
| 株高 Plant height/cm | 112.3±5.2 | 98.9±5.5 | 0.0003 |
| 单株有效穗数 No. of panicles per plant | 8.1±1.4 | 6.3±1.8 | 0.0227 |
| 每穗总粒数 No. of spikelets per panicle | 118.0±9.1 | 96.2±7.3 | <0.0001 |
| 每穗实粒数 No. of grains per panicle | 71.1±6.7 | 51.5±5.9 | <0.0001 |
| 千粒重 1000-grain weight /g | 33.5±0.4 | 31.9±0.5 | 0.3067 |
| 单株产量 Grain yield per plant /g | 19.3±2.8 | 10.3±1.4 | <0.0001 |
Table 2 Comparison of main agronomic traits between the mutant and its wild-type parent.
| 性状 Trait | 中恢8015 R8015 | w390 | P |
|---|---|---|---|
| 株高 Plant height/cm | 112.3±5.2 | 98.9±5.5 | 0.0003 |
| 单株有效穗数 No. of panicles per plant | 8.1±1.4 | 6.3±1.8 | 0.0227 |
| 每穗总粒数 No. of spikelets per panicle | 118.0±9.1 | 96.2±7.3 | <0.0001 |
| 每穗实粒数 No. of grains per panicle | 71.1±6.7 | 51.5±5.9 | <0.0001 |
| 千粒重 1000-grain weight /g | 33.5±0.4 | 31.9±0.5 | 0.3067 |
| 单株产量 Grain yield per plant /g | 19.3±2.8 | 10.3±1.4 | <0.0001 |
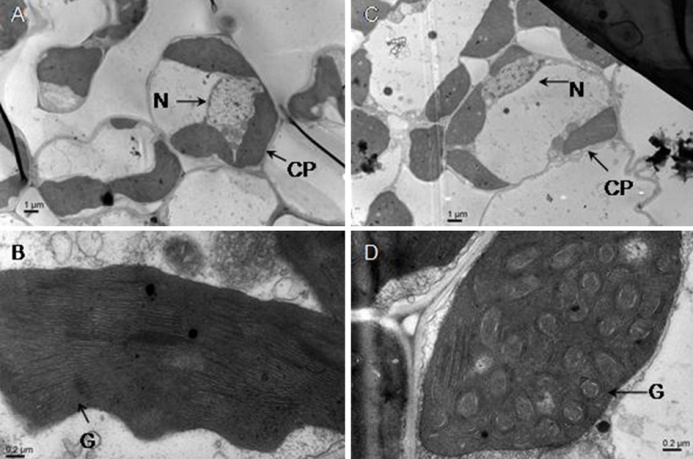
Fig. 2. Comparison of ultrastructures of chloroplasts in the mesophyll cells between the mutant and its wild-type parent.(A and B, Mesophyll cell and chloroplast of wild-type, respectively; C and D, Mesophyll cell and chloroplast of mutant, respectively. N, Nucleus; CP, Chloroplast; G, Grana.)
| 引物名称 Primer name | 上游引物 Forward primer (5'-3') | 下游引物 Reverse primer (5'-3') | 退火温度 Anealing temperature/℃ |
|---|---|---|---|
| D2 | CAGCCTCTCTAATTGACTCTC | CCAAGTAGAATGGCCAAATGT | 55 |
| D3 | GCATCAAACTATAATAACTGACT | AGGAAAGTAAACAAGGCCTTA | 55 |
| D4 | ACACATATGTAGAGTATTATCCG | GCAAGCTGTCTACACGGTT | 55 |
| D8 | AACTCTATTGTTTATGGTGG | TAATTCCGAATTTCAGCTCT | 55 |
| CAO1-1 | CATGCCTACTTGTGTCACT | ACCGCTCATGTGTACCATC | 55 |
| CAO1-2 | GGCGTATTCCAAACCTATTCG | TTAGAATAAAATACGGTGTGCT | 55 |
| CAO1-3 | ATTTCCTACTACCCGAAGCTG | GACTATAGTATGCGGTTACCTT | 55 |
| CAO1-4 | GATTACAATTGCATCCCGTGA | ATGCCATCCACAAAGATGCTC | 55 |
| CAO1-5 | CAGAAGAAGTCCATGCTCCA | CCGGTTCGATATCCAGTATTGC | 55 |
| CAO1-6 | AGATGATACTCTAGTTTCCGACA | AATTAGACAAAACCACCCTCG | 60 |
Table 3 Markers used for fine mapping and sequencing.
| 引物名称 Primer name | 上游引物 Forward primer (5'-3') | 下游引物 Reverse primer (5'-3') | 退火温度 Anealing temperature/℃ |
|---|---|---|---|
| D2 | CAGCCTCTCTAATTGACTCTC | CCAAGTAGAATGGCCAAATGT | 55 |
| D3 | GCATCAAACTATAATAACTGACT | AGGAAAGTAAACAAGGCCTTA | 55 |
| D4 | ACACATATGTAGAGTATTATCCG | GCAAGCTGTCTACACGGTT | 55 |
| D8 | AACTCTATTGTTTATGGTGG | TAATTCCGAATTTCAGCTCT | 55 |
| CAO1-1 | CATGCCTACTTGTGTCACT | ACCGCTCATGTGTACCATC | 55 |
| CAO1-2 | GGCGTATTCCAAACCTATTCG | TTAGAATAAAATACGGTGTGCT | 55 |
| CAO1-3 | ATTTCCTACTACCCGAAGCTG | GACTATAGTATGCGGTTACCTT | 55 |
| CAO1-4 | GATTACAATTGCATCCCGTGA | ATGCCATCCACAAAGATGCTC | 55 |
| CAO1-5 | CAGAAGAAGTCCATGCTCCA | CCGGTTCGATATCCAGTATTGC | 55 |
| CAO1-6 | AGATGATACTCTAGTTTCCGACA | AATTAGACAAAACCACCCTCG | 60 |
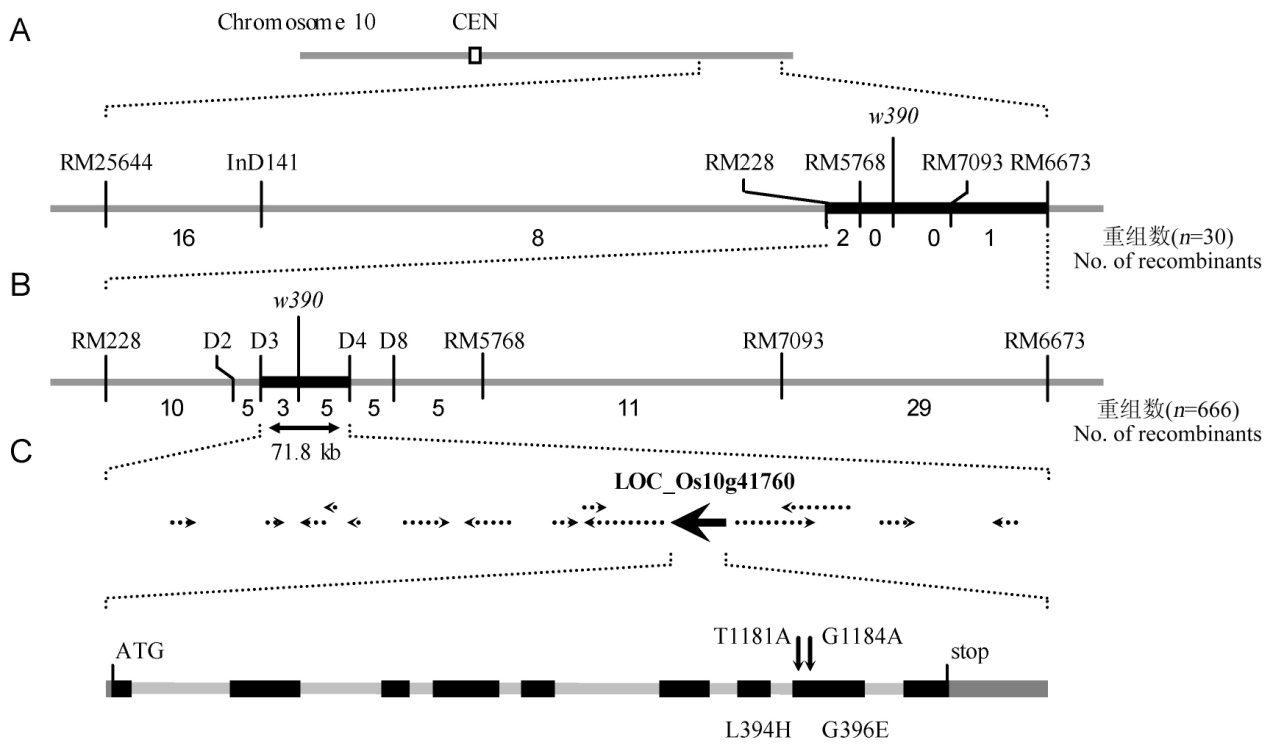
Fig. 3. Fine mapping and positional cloning of the mutant gene.(A, The mutant locus was mapped on the chromosome 10 between RM228 and RM6673. B, The locus was narrowed to a 71.8 kb genomic region between InDel marker D3 and D4. C, Sequence analysis revealed that the mutant carried two nucleotide substitutions in the eighth exon of LOC_Os10g41760, which led to the substitution of leucine and glycine for histidine and glutamic acid, respectively.)
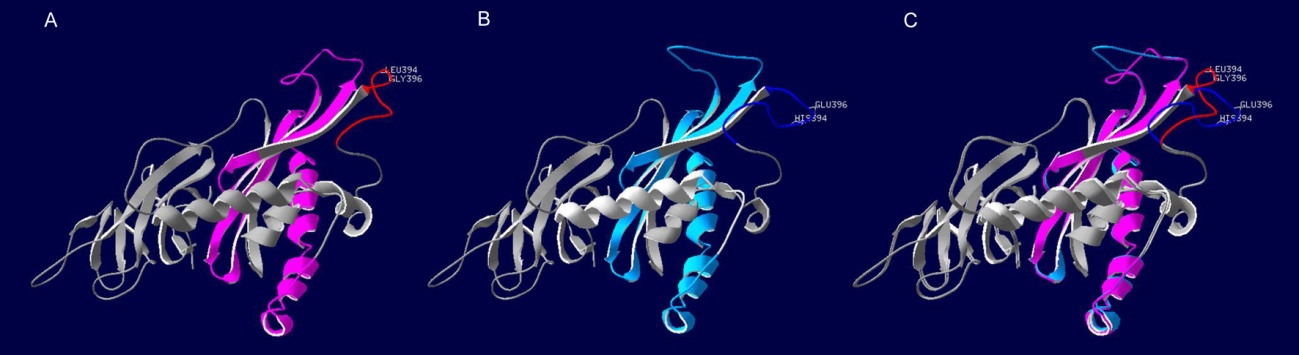
Fig. 4. Comparison of 3D protein structure model of CAO1 between wild-type and the mutant.(A, Wild-type; B, Mutant; C, Superposition of the mutant and its wild-type. The red and dark blue lines represent the structure haboring the mutations in the mutant and its wild-type, respectively. The purple and light blue regions represent the pheophorbide a oxygenase(PaO) domain in the mutant and its wild-type, respectively.)
| [1] | Samuel I B.Green genes gleaned.Trends Plant Sci, 2005, 10(7): 309-312. |
| [2] | Kusaba M, Ito H, Morita R, et al.Rice NON-YELLOW COLORING1 is involved in light-harvesting complex II and grana degradation during leaf senescence.Plant Cell, 2007, 19: 1362-1375. |
| [3] | Wang P R, Gao J X, Wan C M, et al.Divinyl chlorophyll (ide) a can be converted to monovinyl chlorophyll(ide) a by a divinyl reductase in rice.Plant Physiol, 2010, 153: 994-1003. |
| [4] | Yamatani H, Sato Y, Masuda Y, et al.NYC4, the rice ortholog of Arabidopsis THF1, is involved in the degradation of chlorophyll-protein complexes during leaf senescence.Plant J, 2013, 74: 652-662. |
| [5] | Sugimoto H, Kusumi K, Tozawa Y, et al.The virescent-2 mutation inhibits translation of plastid transcripts for the plastid genetic system at an early stage of chloroplast differentiation.Plant Cell Physiol, 2004, 45(8): 985-996. |
| [6] | Kusumi K, Sakata C, Nakamura T, et al.A plastid protein NUS1 is essential for build-up of the genetic system for early chloroplast development under cold stress conditions.Plant J, 2011, 68(6): 1039-1050. |
| [7] | Su N, Hu M L, Wu D X, et al.Disruption of a rice pentatricopeptide repeat protein causes a seedling-specific albino phenotype and its utilization to enhance seed purity in hybrid rice production.Plant Physiol, 2012, 159(1): 227-238. |
| [8] | 邓晓娟, 张海清, 王悦, 等. 水稻叶色突变基因研究进展. 杂交水稻, 2012, 27(5): 9-14. |
| [9] | 李贤勇, 王楚桃, 李顺武, 等. 一个水稻高叶绿素含量基因的发现. 西南农业学报, 2002, 15(4): 122-123. |
| [10] | Juan M P, María C S, Kerstin K, et al.Rice NTRC is a high-efficiency redox system for chloroplast protection against oxidative damage.Plant Cell, 2006, 18(9): 2356-2368. |
| [11] | Miyoshi K, Ito Y, Serizawa A, et al.OsHAP3 genes regulate chloroplast biogenesis in rice.Plant J, 2003, 36(4): 532-540. |
| [12] | Dong H, Fei G L, Wu C Y, et al.A rice virescent-yellow leaf mutant reveals new insights into the role and assembly of plastid caseinolytic protease in higher plants.Plant Physiol, 2013, 162(4): 1867-1880. |
| [13] | Zhao C F, Xu J M, Chen Y, et al.Molecular cloning and characterization of OsCHR4, a rice chromatin-remodeling factor required for early chloroplast development in adaxial mesophyll.Planta, 2012, 236(4): 1165-1176. |
| [14] | Liu W Z, Fu Y P, Hu G C, et al.Identification and fine mapping of a thermo-sensitive chlorophyll deficient mutant in rice (Oryza sativa L.).Planta, 2007, 226(3): 785-795. |
| [15] | Tsugane K, Maekawa M, Takagi K, et al.An active DNA transposon nDart causing leaf variegation and mutable dwarfism and its related elements in rice.Plant J, 2006, 45(1): 46-57. |
| [16] | Gothandam K M, Kim E S, Cho H, et al.OsPPR1, a pentatricopeptide repeat protein of rice is essential,for the chloroplast biogenesis.Plant Mol Biol, 2005, 58: 421-433. |
| [17] | Goh C H, Jung K H, Roberts S K, et al.Mitochondria provide the main source of cytosolic ATP for activation of outward-rectifying K+ channels in mesophyll protoplast of chlorophyll-deficient mutant rice (OsCHLH) seedlings.J Biol Chem, 2004, 279: 6874-6882. |
| [18] | Zhang H T, Li J J, Yoo J H, et al.Rice Chlorina-1 and Chlorina-9 encode ChlD and ChlI subunits of Mg-chelatase, a key enzyme for chlorophyll synthesis and chloroplast development.Plant Mol Biol, 2006, 62(3): 325-337. |
| [19] | Sakuraba Y, Rahman M L, Cho S H, et al.The rice faded green leaf locus encodes protochlorophyllide oxidoreductase B and is essential for chlorophyll synthesis under high light conditions.Plant J, 2013, 74(1): 122-133. |
| [20] | Lee S, Kim J H, Yoo E S, et al.Differential regulation of chlorophyll a oxygenase genes in rice.Plant Mol Biol, 2005, 57(6): 805-818. |
| [21] | 杨海莲, 刘敏, 郭旻, 等. 一个水稻黄绿叶突变体ygl10的遗传分析和基因定位. 中国水稻科学, 2014, 28(1): 41-48. |
| [22] | Li J, Long Y, Qi G N, et al.The Os-AKT1 channel Is critical for K+ uptake in rice roots and is modulated by the rice CBL1-CIPK23 complex.Plant Cell, 2014, 80: 300-307. |
| [23] | Wu Z M, Zhang X, He B, et al.A chlorophyll-deficient rice mutant with impaired chlorophyllide esterification in chlorophyll biosynthesis.Plant Physiol, 2007, 145(1): 29-40. |
| [24] | Morita R, Sato Y, Masuda Y, et al.Defect in non-yellow coloring 3, an α/β hydrolase-fold family protein, causes a stay-green phenotype during leaf senescence in rice.Plant J, 2009, 59(6): 940-952. |
| [25] | Jiang H W, Li M R, Liang N T, et al.Molecular cloning and function analysis of the stay green gene in rice.Plant J, 2007, 52(2): 197-209. |
| [26] | Fang J, Chai C L, Qian Q, et al.Mutations of genes in synthesis of the carotenoid precursors of ABA lead to pre-harvest sprouting and photo-oxidation in rice.Plant J, 2008, 54(2): 177-189. |
| [27] | Li Z, Zhang Y X, Liu L, et al.Fine mapping of the lesion mimic and early senescence 1 (lmes1) in rice (Oryza sativa L.).Plant Physiol Biochem, 2014, 80: 300-307. |
| [28] | SAS Institute Inc, SAS/STAT User’s Guide. SAS Institute, Cary, NC, USA, 1999. |
| [29] | Rogers S O, Bendich A J.Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues.Plant Mol Biol, 1985, 5: 69-76. |
| [30] | Orjuela J, Garavito A, Bouniol M, et al.A universal core genetic map for rice.Theor Appl Genet, 2010, 120: 563-572. |
| [31] | Rychlik W.Oligo Primer Analysis Software Version 7.0. 2nd ed. Molecular Biology Insights, Inc, Cascade, CO, 2008. |
| [32] | 刘胜, 魏祥进, 邵高能, 等. 一个水稻“斑马叶”叶色突变体基因zebraleaf2(zl2)的图位克隆. 中国水稻科学, 2013, 27(3): 231-239. |
| [33] | Yamasato A, Nagata N, Tanaka R, et al.The N-Terminal domain of chlorophyllide a oxygenase confers protein instability in response to chlorophyll b accumulation in Arabidopsis.Plant Cell, 2005, 17: 1585-1597. |
| [34] | Harrison M A, Nemson J A, Melis A.Assembly and composition of the chlorophyll a-b light-harvesting complex of barley (Hordeum vulgare L.): Immunochemical analysis of chlorophyll b-less and chlorophyll b-deficient mutants.Photosyn Res, 1993, 38: 141-151. |
| [35] | Staehelin L A.Chloroplast structure and supramolecular organization of photosynthetic membranes// Pirson A, Zimmermann M H. Encyclopedia of Plant Physiology. Berlin: Germany:Springer-Verlag,1986: 1-84. |
| [36] | Rüdiger W.Biosynthesis of chlorophyll b and the chlorophyll cycle.Photosyn Res, 2002, 74: 187-193. |
| [1] | HOU Xiaoqin, WANG Ying, YU Bei, FU Weimeng, FENG Baohua, SHEN Yichao, XIE Hangjun, WANG Huanran, XU Yongqiang, WU Zhihai, WANG Jianjun, TAO Longxing, FU Guanfu. Mechanisms Behind the Role of Potassium Fulvic Acid in Enhancing Salt Tolerance in Rice Seedlings [J]. Chinese Journal OF Rice Science, 2024, 38(4): 409-421. |
| [2] | GAO Junru, QUAN Hongyu, YUAN Liuzhen, LI Qinying, QIAO Lei, LI Wenqiang. Map-based Cloning and Functional Analysis of a New Allele of D1, a Gene Controlling Plant Height in Rice (Oryza sativa L.) [J]. Chinese Journal OF Rice Science, 2024, 38(2): 140-149. |
| [3] | XU Huan, ZHOU Tao, SUN Yue, WANG Mumei, YANG Yachun, MA Hui, LI Hao, XU Dawei, ZHOU Hai, YANG Jianbo, NI Jinlong. Characterization and Gene Mapping of a Glume Lesion Mimic Mutant glmm1 in Rice [J]. Chinese Journal OF Rice Science, 2023, 37(5): 497-506. |
| [4] | TANG Jie, LONG Tuan, WU Chunyu, LI Xinpeng, ZENG Xiang, WU Yongzhong, HUANG Peijin. Identification and Gene Mapping of a New Photo-thermo-sensitive Male Sterile Mutant tms3650 in Rice [J]. Chinese Journal OF Rice Science, 2023, 37(1): 45-54. |
| [5] | SUN Zhiguang, DAI Huimin, CHEN Tingmu, LI Jingfang, CHI Ming, ZHOU Zhenling, LIU Yan, LIU Jinbo, XU Bo, XING Yungao, YANG Bo, LI Jian, LU Baiguan, FANG Zhaowei, WANG Baoxiang, XU Dayong. Phenotypic Identification and Gene Mapping of a Lesion Mimic Mutant lmm7 in Rice [J]. Chinese Journal OF Rice Science, 2022, 36(4): 357-366. |
| [6] | QIU Linlin, LIU Qiao, FU Yaping, LIU Wenzhen, HU Guocheng, ZHAI Yufeng, PANG Bo, WANG Dekai. Identification and Gene Cloning of DSP2 in Rice (Oryza sativa L.) [J]. Chinese Journal OF Rice Science, 2022, 36(2): 150-158. |
| [7] | LIANG Cheng, XIANG Xunchao, ZHANG Ouling, YOU Hui, XU Liang, CHEN Yongjun. Analyses on Agronomic Traits and Genetic Characteristics of Two New Plant-architecture Lines in Rice [J]. Chinese Journal OF Rice Science, 2022, 36(2): 171-180. |
| [8] | YANG Jinyu, BAI Chen, DING Xiaohui, SHEN Hongfang, WANG Lei, YING Jiezheng, E Zhiguo. Genetic Analysis and Gene Mapping of a Male Sterile Mutant ms7 in Rice [J]. Chinese Journal OF Rice Science, 2022, 36(1): 27-34. |
| [9] | Chen WANG, Beifang WANG, Yingxin ZHANG, Yongrun CAO, Yue ZHANG, Min JIANG, Kangji BIAN, Xiaohui ZHANG, Qunen LIU. Identification and Gene Mapping of a Lesion Mimic Mutant lm8015-2 in Rice [J]. Chinese Journal OF Rice Science, 2021, 35(4): 352-358. |
| [10] | Yujun ZHU, Ziwei ZUO, Zhenhua ZHANG, Yeyang FAN. A New Approach for Fine-mapping and Map-based Cloning of Minor-Effect QTL in Rice [J]. Chinese Journal OF Rice Science, 2021, 35(4): 407-414. |
| [11] | Wei LIU, Zhanhua LU, Dongbai LU, Xiaofei WANG, Shiguang WANG, Jia XUE, Xiuying HE. Location and Candidate Gene Analysis of Rice Clustered Spikelets Gene OsCL6 [J]. Chinese Journal OF Rice Science, 2021, 35(3): 238-248. |
| [12] | Xuemei DENG, Peng HU, Yueying WANG, Yi WEN, Yiqing TAN, Hao WU, Kaixiong WU, Junge WANG, Linlin HOU, Lixin ZHU, Li ZHU, Guang CHEN, Dali ZENG, Guangheng ZHANG, Longbiao GUO, Zhenyu GAO, Deyong REN, Qian QIAN, Jiang HU. Identification and Fine Mapping of a Grain Width Mutant gw4 in Rice [J]. Chinese Journal OF Rice Science, 2021, 35(3): 259-268. |
| [13] | Shufang LIU, Liying DONG, Xundong LI, Wumin ZHOU, Qinzhong YANG. Different Reactions of Rice Monogenic Line IRBL9-W Harboring Pi9 Gene to Magnaporthe oryzae Containing AvrPi9 During Seedling and Adult-plant Stages [J]. Chinese Journal OF Rice Science, 2021, 35(3): 303-310. |
| [14] | Yali ZHENG, Linchuang YU, Xiaoxiao AN, Xinle CHENG, Lijun REN, Zilong SU, Xiaoya ZHENG, Tao LAN. Identification of a Knockout Mutant of OsWOX3B Gene in Rice (Oryza sativa L.) [J]. Chinese Journal OF Rice Science, 2021, 35(2): 112-120. |
| [15] | Zhonghao WANG, Yan HE, Xiaobo ZHANG, Xia XU, WUJianli, SHIYongfeng. Genetic Analysis and Gene Mapping of a Virescent and Panicle AbortionMutant vpa1in Rice [J]. Chinese Journal OF Rice Science, 2021, 35(1): 19-26. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||