
Chinese Journal OF Rice Science ›› 2015, Vol. 29 ›› Issue (3): 223-231.DOI: 10.3969/j.issn.1001G7216.2015.03.001
• ResearchPaper • Next Articles
Xiu-mei WANG, Yue-yang LIANG, Ling LI, Chang-wei GONG, Hai-peng WANG, Xiao-xi HUANG, Shuang-cheng LI, Qi-ming DENG, Jun ZHU, Ai-ping ZHENG, Ping LI, Shi-quan WANG*( )
)
Received:2014-11-15
Revised:2014-12-23
Online:2015-05-10
Published:2015-05-10
Contact:
Shi-quan WANG
王秀梅, 梁越洋, 李玲, 贡常委, 王海鹏, 黄晓西, 李双成, 邓其明, 朱军, 郑爱萍, 李平, 王世全*( )
)
通讯作者:
王世全
CLC Number:
Xiu-mei WANG, Yue-yang LIANG, Ling LI, Chang-wei GONG, Hai-peng WANG, Xiao-xi HUANG, Shuang-cheng LI, Qi-ming DENG, Jun ZHU, Ai-ping ZHENG, Ping LI, Shi-quan WANG. OsMAX1a and OsMAX1e, Involved in the Biosynthesis of Strigolactones,Regulate Rice Tillering[J]. Chinese Journal OF Rice Science, 2015, 29(3): 223-231.
王秀梅, 梁越洋, 李玲, 贡常委, 王海鹏, 黄晓西, 李双成, 邓其明, 朱军, 郑爱萍, 李平, 王世全. OsMAX1a, OsMAX1e通过参与独角金内酯的合成调控水稻分蘖[J]. 中国水稻科学, 2015, 29(3): 223-231.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.3969/j.issn.1001G7216.2015.03.001
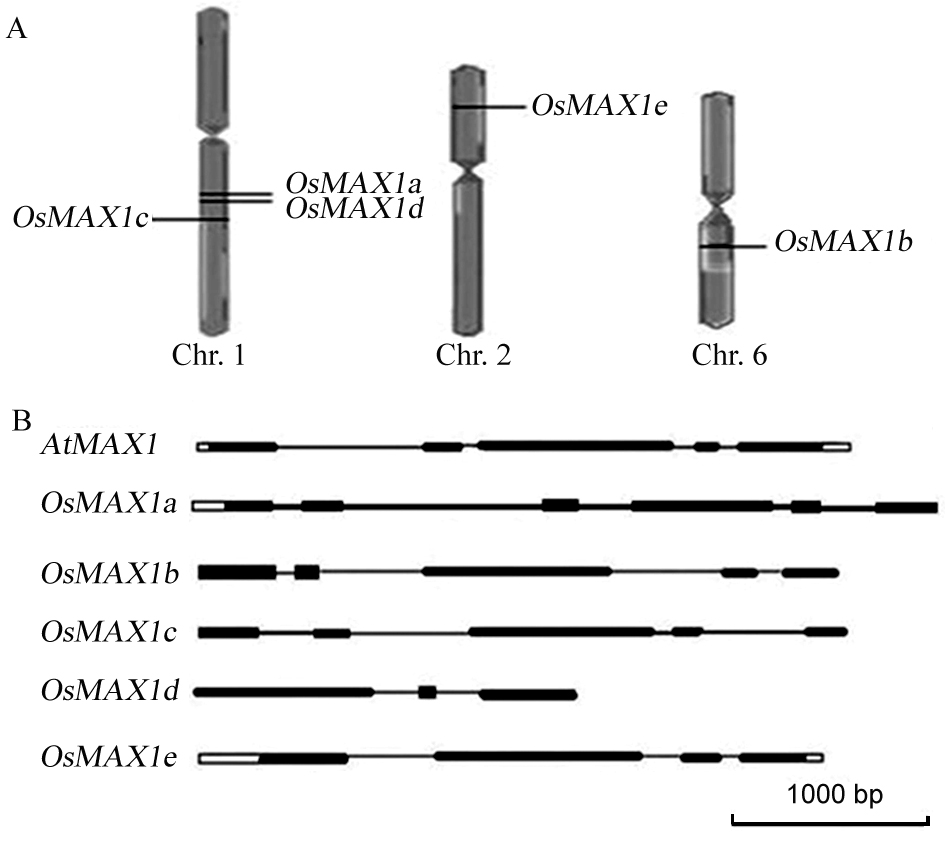
Fig. 1. Chromosomal location and gene structure of rice OsMAX1 candidate genes.(In Fig. B, black boxes indicate exon and the connecting lines indicate intron.)
| 基因 Gene | 登录号 ID | 染色体 Chromosome | 核苷酸大小 Nucleotide size / aa |
|---|---|---|---|
| OsMAX1a | LOC_Os01g50530 | 1 | 412 |
| LOC_Os01g50520 | 1 | ||
| OsMAX1b | LOC_Os06g36920 | 6 | 549 |
| OsMAX1c | LOC_Os01g50590 | 1 | 517 |
| OsMAX1d | LOC_Os01g50580 | 1 | 385 |
| OsMAX1e | LOC_Os02g12890 | 2 | 548 |
Table 1 The OsMAX1 candidate genes in rice.
| 基因 Gene | 登录号 ID | 染色体 Chromosome | 核苷酸大小 Nucleotide size / aa |
|---|---|---|---|
| OsMAX1a | LOC_Os01g50530 | 1 | 412 |
| LOC_Os01g50520 | 1 | ||
| OsMAX1b | LOC_Os06g36920 | 6 | 549 |
| OsMAX1c | LOC_Os01g50590 | 1 | 517 |
| OsMAX1d | LOC_Os01g50580 | 1 | 385 |
| OsMAX1e | LOC_Os02g12890 | 2 | 548 |
| 引物 Marker | 上游引物 Sence primer | 下游引物 Anti-sence primer |
|---|---|---|
| RNAi-OsMAX1CON-F | 5'-GCGTCGACGTTCTCAAGAGGCTTCGCTG-3' | 5'-CGAGCTCGTTCTCAAGAGGCTTCGCTG-3' |
| RNAi-OsMAX1CON-R | 5'-CCCAAGCTTCAAGGTAGGGGAATTTGGTCT-3' | 5'-CGGAATTCCAAGGTAGGGGAATTTGGTCT-3' |
| RNAi-OsMAX1a-F | 5'-GCGTCGAC TCCTGAAGGAAGCGATGAGA-3' | 5'-CGAGCTC TCCTGAAGGAAGCGATGAGA-3' |
| RNAi-OsMAX1a-R | 5'-CCCAAGCTTCACCTCTGGCTCTGGGAAAT-3' | 5'-CGGAATTCCACCTCTGGCTCTGGGAAAT-3' |
| RNAi-OsMAX1e-F | 5'-GCGTCGACGTCCAGCTCGCTGTCCACCA-3' | 5'-TCCCCCGGGGTCCAGCTCGCTGTCCACCA-3' |
| RNAi-OsMAX1e-R | 5'-CCCAAGCTTGAGACGAGGTGGAAGCGCATG-3' | 5'-CGGAATTCGAGACGAGGTGGAAGCGCATG-3' |
Table 2 Sequence of RNAi markers used in the study.
| 引物 Marker | 上游引物 Sence primer | 下游引物 Anti-sence primer |
|---|---|---|
| RNAi-OsMAX1CON-F | 5'-GCGTCGACGTTCTCAAGAGGCTTCGCTG-3' | 5'-CGAGCTCGTTCTCAAGAGGCTTCGCTG-3' |
| RNAi-OsMAX1CON-R | 5'-CCCAAGCTTCAAGGTAGGGGAATTTGGTCT-3' | 5'-CGGAATTCCAAGGTAGGGGAATTTGGTCT-3' |
| RNAi-OsMAX1a-F | 5'-GCGTCGAC TCCTGAAGGAAGCGATGAGA-3' | 5'-CGAGCTC TCCTGAAGGAAGCGATGAGA-3' |
| RNAi-OsMAX1a-R | 5'-CCCAAGCTTCACCTCTGGCTCTGGGAAAT-3' | 5'-CGGAATTCCACCTCTGGCTCTGGGAAAT-3' |
| RNAi-OsMAX1e-F | 5'-GCGTCGACGTCCAGCTCGCTGTCCACCA-3' | 5'-TCCCCCGGGGTCCAGCTCGCTGTCCACCA-3' |
| RNAi-OsMAX1e-R | 5'-CCCAAGCTTGAGACGAGGTGGAAGCGCATG-3' | 5'-CGGAATTCGAGACGAGGTGGAAGCGCATG-3' |
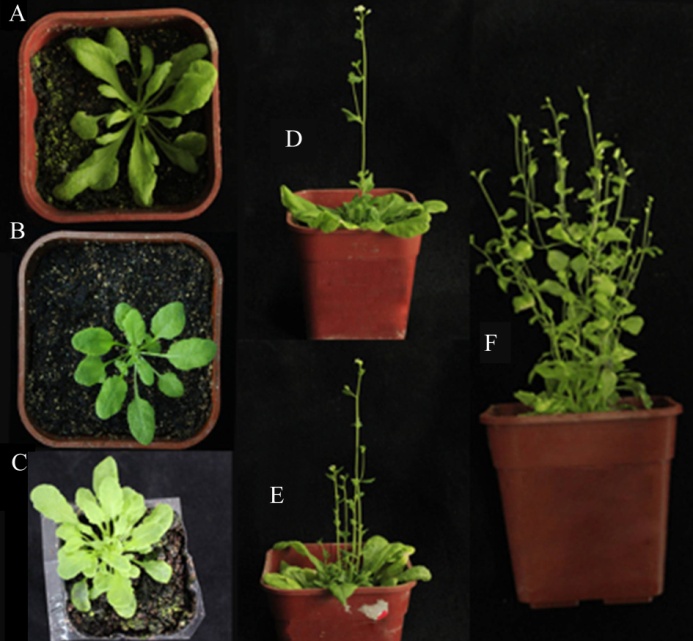

Fig.3. Overexpression of OsMAX1a and OsMAX1e gene in Arabidopsis max1. (A and D, Over-expression of OsMAX1a in Arabidopsis max1; B and E, Over-expression of OsMAX1e in Arabidopsis max1; C and F, Arabidopsis max1.)
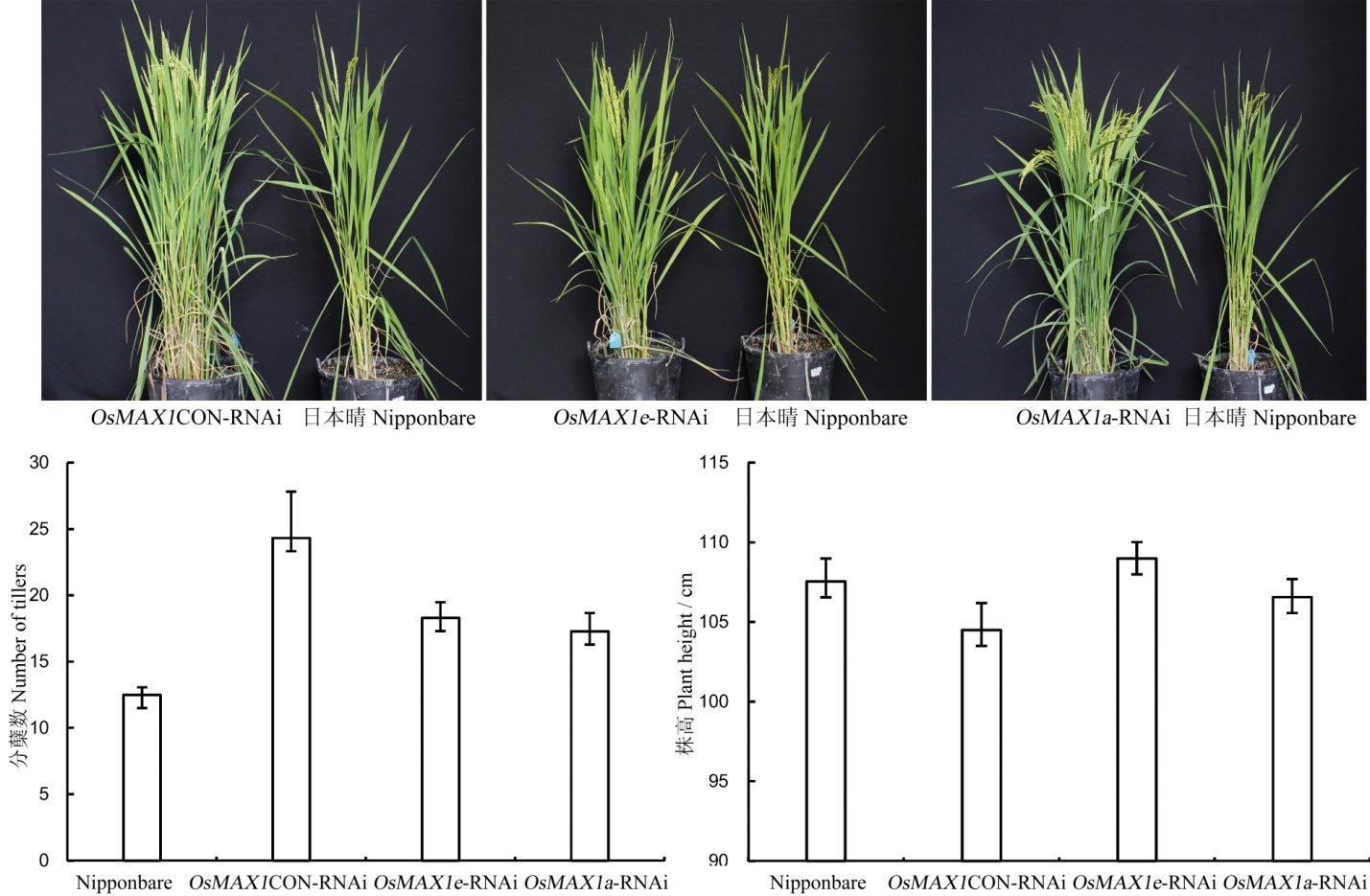
Fig. 5. Tiller number and height comparison of Nipponbare, OsMAX1a and OsMAX1e gene interference in rice.(OsMAX1CON-RNAi, The conserved region interference of OsMAX1 gene.)
| [1] | Ruyter-Spira C, Al-Babili S, van der Krol S, et al. The biology of strigolactones.Trends Plant Sci, 2013, 18: 72-83 . |
| [2] | Wang Y H, Li J Y.Branching in rice.Plant Biol, 2011, 14: 94-99. |
| [3] | Domagalska M A, Leyser O.Signal integration in the control of shoot branching.Cell Biol, 2011(2): 211-221. |
| [4] | Cardoso C, Ruyter-Spira C, Bouwmeester H J, et al.Strigolactones and root infestation by plant-parasitic Striga, Orobanche and Phelipanche spp.Plant Sci, 2011, 180(3): 414-420. |
| [5] | Dun E A, Brewer P B, Beveridge C A, et al.Strigolactones: Discovery of the elusive shoot branching hormone.Trends Plant Sci, 2009, 14: 364-372. |
| [6] | Stirnberg P, van De Sande K, Leyser H M. MAX1 and MAX2 control shoot lateral branching in Arabidopsis.Development, 2002, 129: 1131-1141. |
| [7] | Umehara M, Hanada A, Yoshida S, et al.Inhibition of shoot branching by new terpenoid plant hormones.Nature, 2008, 455: 195-200. |
| [8] | Ruyter-Spira C, Al-Babili S, van der Krol, et al. The biology of strigolactones.Trends Plant Sci, 2013, 18: 72-83. |
| [9] | Woo H R, Chung K M, Park J H, et al.ORE9, an F-box protein that regulates leaf senescence in Arabidopsis.Plant Cell, 2001, 13: 1779-1790. |
| [10] | Gomez-Roldan, Fermas V S, Brewer P B, et al.Strigolactone inhibition of shoot branching.Nature, 2008, 455: 189-194. |
| [11] | Snowden K C, Simkin A J, Janssen B J, et al.The decreased apical dominance1/petunia hybrid CAROTENOID CLEAVAGE DIOXYGENASE 8 gene affects branch production and plays a role in leaf senescence, root growth, and flower development.Plant Cell, 2005, 17: 746-759. |
| [12] | Cook C G, Whichard L P, Turner B, et al.Germination of witchweed (Striga lutea Lour.): Isolation and properties of a potent stimulant.Science, 1966, 154: 1189-1190. |
| [13] | Akiyama K, Matsuzaki K, Hayashi H, et al.Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi.Nature, 2005, 435: 824-827. |
| [14] | Umehara M, Hanada A, Yoshida S, et al.Inhibition of shoot branching by new terpenoid plant hormones.Nature, 2008, 455: 195-200. |
| [15] | Kohlen W, Charnikhova T, Lammers M, et al.Strigolactones are transported through the xylem and play a key role in shoot architectural response to phosphate deficiency in nonarbuscular mycorrhizal host Arabidopsis.Plant Physiol, 2011, 155(2): 974-987. |
| [16] | Zhou F, Lin Q B, Zhu L H, et al.D14-SCFD3-dependent degradation of D53 regulates strigolactone signalling.Nature, 2013, 504: 7480. |
| [17] | Zou J, Zhang S Y, Zhang W P, et al.The rice HIGH-TILLERING DWARF1 encoding an ortholog of Arabidopsis MAX3 is required for negative regulation of the outgrowth of axillary buds.Plant J, 2006, 48(5): 687-698. |
| [18] | Arite T, Umehara M, Ishikawa S, et al.d14, a strigolactone-insensitive mutant of rice, shows an accelerated outgrowth of tillers.Plant Cell Physiol, 2009, 50: 1416-1424. |
| [19] | Lin H, Wang R, Qian Q, et al.DWARF27, an iron-containing protein required for the biosynthesis of strigolactones, regulates rice tiller bud outgrowth.Plant Cell, 2009, 21: 1512-1525. |
| [20] | Arite T, Iwata H, Ohshima K, et al.DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral budoutgrowth in rice.Plant J, 2007, 51: 1019-1029. |
| [21] | Jiang L, Liu X, Xiong G S, et al.DWARF 53 acts as a repressor of strigolactone signalling in rice.Nature, 2013, 504(7480): 401-405. |
| [22] | Booker J, Auldridge M, Wills S, et al.MAX3/CCD7 is a carotenoid cleavage dioxygenase required for the synthesis of a novel plant signaling molecule.Curr Biol, 2004, 14: 1232-1238. |
| [23] | Sorefan K, Booker J, Haurogné K, et al.MAX4 and RMS1 are orthologous dioxygenase-like genes that regulate shoot branching in Arabidopsis and pea.Genes Dev, 2003, 17: 1469-1474. |
| [24] | Beveridge C A, Symons G M, Turnbull C G.Long-distance signaling and a mutation analysis of branching in pea.Plant Growth Regul, 2000, 32: 193-203. |
| [25] | Hamiaux C, Drummond R S, Janssen B J, et al.DAD2 is an a/b hydrolase likely to be involved in the perception of the plant branching hormone, strigolactone.Curr Biol, 2012, 22: 2032-2036. |
| [26] | Huang J Q, Wei Z M, An H L, et al.Agrobacterium tumefaciens-mediated transformation of rice with the spider insecticidalgene conferring resistance to leaffolder and striped stem borer.Cell Res, 2001, 11: 149-155. |
| [27] | Hiei Y, Ohta S, Komari T, et al.Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA.Plant J, 1994, 6: 271-282. |
| [28] | 许红梅, 张立军, 刘淳. 农杆菌蘸花法侵染拟南芥的研究. 北方园艺, 2010, 14: 143-146. |
| [29] | Yoshid A S, Forno D A, Cock J H, et al.Laboratory manual for physiological studies of rice. 3rd ed. Manila, Philippines: IRRI, 1976: 61-64. |
| [30] | Turnbull C G N, Booker J P, Leyser H M O. Micrografting techniques for testing long-distance signalling in Arabidopsis.Plant J, 2002, 32: 255-262. |
| [31] | Booker J, Sieberer T, Wright W, et al.MAX1 encodes a cytochrome P450 family member that acts downstream of MAX3/4 to produce a carotenoid-derived branch-inhibiting hormone.Dev Cell, 2005, 8: 443-449. |
| [32] | Alder A, Holdermann I, Beyer P, et al.Carotenoid oxygenases involved in plant branching catalyse a highly specific conserved apocarotenoid cleavage reaction.Biochem J, 2008, 416: 289-296. |
| [33] | Alder A, Jamil M, Marzorati M, et al.The path from b-carotene to carlactone, astrigolactone-like plant hormone.Science, 2012, 335: 1348-1351. |
| [34] | Drummond R S, Sheehan H, Simons J L, et al.The expression of petunia strigolactone pathway genesis altered as part of the endogenous developmental program.Front Plant Sci, 2012, 2: 115. |
| [35] | Yoneyama K, Takeuchi T, Sekimoto H, et al.Phosphorus deficiency in red clover promotes exudation of orobanchol, the signal for mycorrhizal symbionts and germination stimulant for root parasites.Planta, 2007, 225(4): 1031-1038. |
| [1] | HU Jijie, HU Zhihua, ZHANG Junhua, CAO Xiaochuang, JIN Qianyu, ZHANG Zhiyuan, ZHU Lianfeng. Effects of Rhizosphere Saturated Dissolved Oxygen on Photosynthetic and Growth Characteristics of Rice at Tillering Stage [J]. Chinese Journal OF Rice Science, 2024, 38(4): 437-446. |
| [2] | Xianmei WU, Sanfeng LI, Ping HU, Rui HE, Ran JIAO, Yijian MAO, Caolin LU, Juan HU, Han LIN, Rongliang WU, Xudong ZHU, Yuchun RAO, Yuexing WANG. Cloning and Functional Analysis of Rice Tillering Regulatory Gene HTD3 [J]. Chinese Journal OF Rice Science, 2021, 35(6): 535-542. |
| [3] | Minjuan ZHANG, Shuaijun LI, Qiongqiong CHEN, Xiuqing JING, Kunming CHEN, Chunhai SHI, Wenqiang LI. Genetic Mapping and Proteomic Analysis of the Dwarf and Low-tillering Mutant dlt3 in Rice [J]. Chinese Journal OF Rice Science, 2018, 32(6): 529-537. |
| [4] | Xiao-mei SU, Yu-xing FANG, Yu-long LIU, Feng LIU, Suo-bing ZHANG, Yun-hui ZHANG, Yi-qun BAO. Genetic Analysis and Gene Mapping of a Dwarf and High-tillering Mutant mz3 in Rice (Oryza sativa L.) [J]. Chinese Journal OF Rice Science, 2016, 30(5): 479-486. |
| [5] | Yun-gao HU, Guo-tao YANG, Lian-an GUO, Peng QIN, Yong-jun CHEN, Shi-gui LI. Genetic Analysis and Mapping of a Dwarf and High-tillering Mutant bf370 in Rice [J]. Chinese Journal OF Rice Science, 2015, 29(4): 357-362. |
| [6] | WANG Tao1, YUAN Shoujiang2, YIN Liang2, ZHAO Jinfeng1, WAN Jianmin1,3, LI Xueyong1,*. Positional Cloning and Expression Analysis of the Gene Responsible for the Hightillering Dwarf Phenotype in the indica Rice Mutant gsor23 [J]. Chinese Journal of Rice Science, 2013, 27(1): 1-8. |
| [7] | ZHANG Yanmei, ZOU Detang* . Association Analysis of Rice Cold Tolerance at Tillering Stage with SSR Markers in japonica Cultivars in Northeast China [J]. Chinese Journal of Rice Science, 2012, 26(4): 423-430. |
| [8] | JIANG Hua ,#,ZHAO Jiang-hong ,#,GUO Long-biao ,JIANG Liang ,XUE Da-wei,ZENG Da-li,QIAN Qian,SUN Guo-chang . QTL Mapping and Epistatic Analysis of High-Order Tillering in Rice (Oryza sativa) [J]. Chinese Journal of Rice Science, 2011, 25(2): 157-162 . |
| [9] | DENG Qi-ming ,WANG Ying-hen ,WANG Shi-quan ,LI Shuang-cheng ,LI Ping. Near Isogonic Lines Establishment and GenomeWide Expression Analysis of a Few Tillering Mutant of Rice [J]. Chinese Journal of Rice Science, 2008, 22(1): 15-22 . |
| [10] | ZHOU Yi ,GUO Shi-wei ,SONG Na ,ZHANG Lei ,SHEN Qi-rong . Effects of Nitrogen Form and Water Stress Interaction on Photosynthesis,Utilization of Water and Nitrogen of Rice Plants at the Tillering Stage [J]. Chinese Journal of Rice Science, 2006, 20(3): 313-318 . |
| [11] | ZHONG Xu-hua,PENG Shao-bing,John E. SHEEHY,LIU Hong-xian <,A style=. Relationship Between Productive Tiller Percentage and Biomass Accumulation in Rice (Oryza sativa L.): A Simulation Approach [J]. Chinese Journal of Rice Science, 2001, 15(2): 107-112 . |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||